1. Types
2. Induction
3. Mechanism
4. Functions
- Homeostasis
- Low nutrients
- Hypoxia
- Defense
Autophagy is molecular mechanism of eukaryotic cells for digesting intracellular content in lysosomes. The word autophagy was coined by C. de Duve in 1963 and stands for "self eating". It has multiple functions. For example, there is always a basal autophagy in the cells for removing faulty or deteriorated cytoplasmic material and it is also a mechanism that balances the metabolic and nutritional states of the cell. Thus, autophagy is involved in energy metabolism, organelle recycling, starting the embryo development, aging, etcetera. In addition, autophagy is activated by any kind of cell stress, like pathogen infection or intracellular malfunctions. Whatever the cause that triggers autophagy, a set of gens, referred as autophagy related genes, are activated. In yeasts, more than 30 genes are being found to be involved in autophagy, of which Atg genes stand out.
1. Types
While the word autophagy is commonly used as synonymous of macroautophagy, there are other types of autophagy in the cell (Figures 1 and 2):
Macroautophagy is a mechanism for enclosing cytoplasmic elements in a large compartment delimited by a double membrane. This compartment is called macroautophagosome and eventually fuses with lysosomes for degrading its content, along with its inner membrane.
Microautophagy involves de inward folding of the lysosome membrane to form small inner lysosomal vesicles, where they are degraded. Cytosolic material is included in this small vesicles. Actually, the formation of vesicles in multivesicular bodies may be regarded as microautophagy.
Chaperone mediated autophagy is the transport of proteins from the cytosol to the interior of the lysosomes helped by transmembrane proteins. Proteins to be imported are selected by other proteins known as chaperones.
Crinophagy is the fusion of vesicles, usually exocytosis vesicles, directly with lysosomes.

2. Induction
Autophagy is induced under cellular stress, like food deprivation, lack of growth factors, infections, oxydative stress, and hypoxia. It can also be induced experimentally, for example, removing amino acids from cell culture serums. A constitutive autophagy is ongoing in all animal cells, so it is a basal and permanent mechanism in these cells. However, plant cells may stop autophagy under environmental or experimental favorable conditions, but start it after cellular stress like low amount of nutrients, high temperature and high salinity, or when some physiological processes are at work like leaf aging, starch store mobilization, pollen germination, etcetera.
3. Mechanism
The molecular mechanism of macroautophy is relatively well-known. Macroautophagy may be selective or non-selective. Selective macroautophagy needs of a previous labelling of the cytosolic material to be degraded and then be recognized by autophagy receptors. This material may be protein aggregates or damaged organelles (mitochondria, endoplasmic reticulum, chloroplasts, and even the nucleus in some cases). During non-selective macroautophagy unspecific cytoplasmic material is enclosed by autophagy membranes, for example during food deprivation.
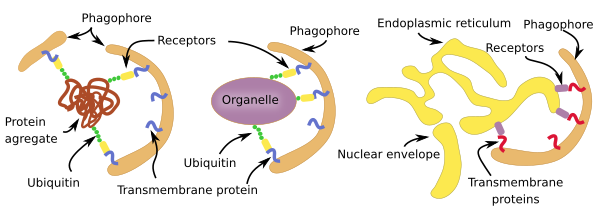
Selective macroautophagy is a molecular mechanism quite similar for most cell structures to be degraded (Figure 3). In animal cells, the labeling is mostly by ubiquitination, that is the addition of the ubiquitin protein as if it was a label. There are receptors in the membranes of the phagophore (see Figure 1) that are able to recognize those linked ubiquitins. For membrane-bound organelles, ubiquitin proteins are added to the cytosolic domain of the membrane proteins. This is the case for peroxin proteins in the peroxisome membrane and for proteins in the mitochondrial outer membrane. However, the endoplasmic reticulum cisterns and tubules and yeast organelles are not selected through ubiquination but by exposing specific proteins in their membranes. There is selective macroautophy capable of degrading intracellular pathogens by ubiquitinating some of their proteins.
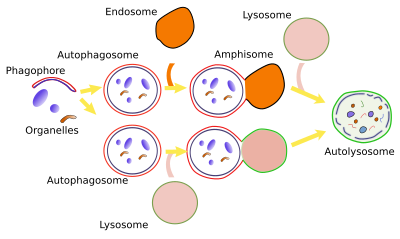
Non-selective macroautophagy, and probably the selective macroautophagy too, begins with the activation of cell endomembranes, which can be part of the endoplasmic reticulum, Golgi apparatus, contact sites between the endoplasmic reticulum and mitochondria. Parts of the plasma membrane may also be activated. The synthesis of lipid PIP3 is the starting point for the activation because the local increase in PIP3 in membranes attracts cytosolic proteins that bend the membrane and detach some fragments of those membrane areas. The fragments join between each other to form the phagophore, also known as isolating membrane. Phagophore membranes enclose portions of cytoplasm and organelles by progressively enlarge the surface and fuse their edges and end up as a closed compartment, the autophagosome, which is a double membrane-bound organelle. Autophagosome, and phagophore, increases in size by acquiring membranes from plasma membrane, ERGIC compartment, Golgi apparatus (there are O-type carbohydrates) and from some vesicles. Autophagosome may directly fuse with a lysosome to form an autophagolysosome, where degradation of the enclosed material happens. Both, late and early endosomes may also send vesicles to the autophagosome or may directly fuse with it. Endosomes provide lysosomal proteins and proton pumps that progressively decrease the pH of the autophagosome, that is, it gets more acid. This progressive pH change transforms the autophagosome into the so-called amphisome. The next step is the fusion of the amphisome with a lysosome to form the autophagolysosome. Whatever the path to get an autophagolysosome, SNARE and Rab proteins need to be in the amphisome or autophagosome to make possible the fusion with the lysosome. In plants, both autophagosome and amphisome fuse with vacuoles for degrading their content because there is no lysosomes.
The number of lysosomes decreases when autophagy is intense after cellular stress. For example, during food deprivation, lysosomes decrease in number after 4 hours, but increase again after 11 hours. Lysosomal population needs to be regenerated again for the cell to be functional again. New lysosomes can emerge from tubules of the autophagolysosome membranes. Cytoskeleton is involved in the formation of these tubules. Clathrin mediated vesicles are generated at the tips of these tubules and, once free in the cytosol, vesicles make their interior progressively more acid until they are mature lysosomes. These vesicles are called protolysosomes because end up as mature lysosomes and the whole process is known as ALR (auphagic lysosome reformation).
4. Functions
In basal state, autophagy helps with cellular homeostasis. However, during cellular stress, the main mission of autophagy is keep the cell alive, at least in the short-term. For that, it degrades huge amount of cellular content with two aims: to decrease the number of cellular compartments that may consume energy and to produce amino acids and other molecules for essential metabolic and cellular processes. However, in the long-term, autophagy is not a desirable mechanism because when the cellular stress is not overcome quickly enough the degradative mechanism ends up damaging so much the cell that it cannot be recovered. Actually, what autophagy is trying to do is gaining time so that cell can better endure and counteract the stress.
Homeostasis
Animal cells always have a basal level of autophagy, even if there is no stress. Another basal degradation pathway in the cell is the ubiquitin-proteosome mechanism, which is specialized in degrading molecules with a short life. However, autophagy is involved in degrading long-live molecules, and, furthermore, it is the only mechanism for degrading organelles such as mitochondria, peroxisomes or endoplasmic reticulum. The basal autophagy may be non-selective and selective. The selective autophagy works as a quality control for removing organelles which are not working properly.
When autophay is inhibited, many types of animal cells shrinks and die. Although plant cells may stop autophagy, some cells need a basal autophagic activity, so that aging and senescent begin when this activity is inhibited, even if there are nutrients enough. It is of notice that autophagy is defective in neurodegenerative pathologies. Some tissular processes like production of surfactant by pneumocytes II, neuromelanin by dopaminergic neurons, and maturation of blood-red cell rely on autophagy. In muscle cells, autophagy removes damaged mitochondria, which is very important for a normal supply of energy. In mammary glands, autophagy is needed for cell death, which is a physiological and normal process.
Low nutrients
In mice, food deprivation induces an increase of autophagy in nearly all tissues, excepting the nervous system. In addition, mutant mice with defective autophagy die after birth, probably because they are not able to counteract the short food deprivation period which happens at this time. Furthermore, they have no basal autophagy. Food deprivation initiates non-selective autophagy. The starting point appears to be the mTORC1 (Target of Rapamycin 1) molecular complex, which is a serine/threonine kinase that responds to energy nutrient levels of the cell and regulates cellular grow and division. mTORC1 may induce catabolism (degradation) or anabolism (synthesis) depending on the cell state. If the amount of amino acids is high, which means good nutrients supply, mTORC1 is activated and inhibits transcription factors responsible for the expression of genes that start autophagy. On the contrary, if intracellular amino acid concentration is low, mTORC1 gets away from lysosomes and becomes inactive. In this situation, genes that stimulate autophagy are activated by transcription factors that enter the nucleus. mTORC1 is associated with lysosome membrane where, somehow, is able to detect the amount of amino acid being degraded inside the lysosome. If lysosomal degradation decreases and the amount of amino acids is low, mTORC1 leaves the lysomal membrane and becomes inactive. Lysosomes without mTORC1 move to the center of the cell where they fuse with autophagolysosomes. Curiously enough, after a few hours of activated autophagy and restored intracellular amino acid levels, mTORC1 is activated again, and one of its roles is to regenerate the lysosomal population from tubules which are formed from autophagosome membranes. These tubules generate vesicles that progressively maturate in pro-lysosomes and later in lysosomes.
Hypoxia
During a short period of myocardial hypoxia there is a temporary decrease of oxygen and nutrients. After that, blood may flow again. During hypoxia (ischemia), macroautophagy is boosted to protect cardiomyocytes from dying by decreasing metabolism and yielding amino acids and other molecules as a source of energy.
Defense against pathogens
Macroautophagy is one of the older cellular defense mechanism and the first defense line against infections by protozoa, bacteria and viruses. Xenophagia is the word for degradation of foreign molecules. There are several levels of actions in the cellular defense response: removing intracellular pathogens and informing the immune system about the infection. TLR (Toll-like receptors) are found in the endosomal membranes. They are able to recognize some DNA sequences from pathogens infecting the cell. The recognition dissociates BCL2/Beclin1, which increases autophagy, and pathogens are conjugated with ubiquitin for their isolation and degradation in autophagosomes. It is of notice that some viruses are able to proliferate inside autophagosomes and get profit from molecules that are being produced.
Bibliography
Choi Y, Bowman JW, Jung JU. 2018. Autophagy during viral infection — a double-edged sword. Nature reviews in microbiology. doi: 10.1038/s41579-018-0003-6.
Eskelinen G-L. 2008. New insights into the mechanisms of macroautophagy in mammalian cells. Internacional review of cell and mollecular biology. 266:207-247.
Gatica D, Lahiri V, Klionsky DJ. 2018. Cargo recognition and degradation by selective autophagy. Nature cell biology 20:233-242.
Klionsky DJ. 2007. Autophagy: from phenomenology to molecular understanding in less than a decade. Nature reviews in molecular and cell biology. doi:10.1038/nrm2245.
Liu Y,Bassham DC. 2012. Autophagy: pathways for self-eating in plant cells. Annual review of plan biology. 63:215-237
Murrow L, Debnath J. 2013. Autophagy as a stress-response and quality-control mechanism: implications for cell injury and human disease.Annual review of pathology. 8:105-137